Expansion Sequencing: Spatially Precise In Situ Transcriptomics in Intact Biological Systems
[Publisher Link] [Local Copy]
Alon S*, Goodwin DR*, Sinha A*, Wassie AT*, Chen F*, Daugharthy ER**, Bando Y, Kajita A, Xue AG, Marrett K, Prior R, Cui Y, Payne AC, Yao CC, Suk HJ, Wang R, Yu CJ, Tillberg P, Reginato P, Pak N, Liu S, Punthambaker S, Iyer EPR, Kohman RE, Miller JA, Lein ES, Lako A, Cullen N, Rodig S, Helvie K, Abravanel DL, Wagle N, Johnson BE, Klughammer J, Slyper M, Waldman J, Jané-Valbuena J, Rozenblatt-Rosen O, Regev A; IMAXT Consortium, Church GM***+, Marblestone AH***, Boyden ES***+ (2021) Expansion Sequencing: Spatially Precise In Situ Transcriptomics in Intact Biological Systems, Science 371(6528):eaax2656. (* equal contribution, ** key contributions to early stages of project, *** equal contribution, +co-corresponding authors)
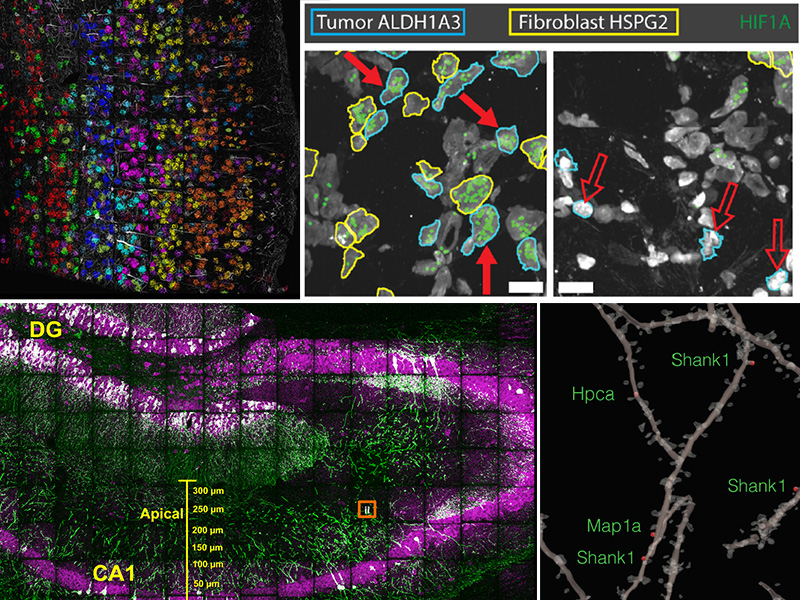
Methods for highly multiplexed RNA imaging are limited in spatial resolution and thus in their ability to localize transcripts to nanoscale and subcellular compartments. We adapt expansion microscopy, which physically expands biological specimens, for long-read untargeted and targeted in situ RNA sequencing. We applied untargeted expansion sequencing (ExSeq) to the mouse brain, which yielded the readout of thousands of genes, including splice variants. Targeted ExSeq yielded nanoscale-resolution maps of RNAs throughout dendrites and spines in the neurons of the mouse hippocampus, revealing patterns across multiple cell types, layer-specific cell types across the mouse visual cortex, and the organization and position-dependent states of tumor and immune cells in a human metastatic breast cancer biopsy. Thus, ExSeq enables highly multiplexed mapping of RNAs from nanoscale to system scale.
Resources associated with this Publication:
[Expansion microscopy: physical magnification with nanoscale precision]